About
I am a multidisciplinary biomedical data scientist with a Ph.D. in Medical Sciences and over six years of postdoctoral experience at
EPFL's Blue Brain Project, where I specialized in computational neuroscience and neuro-glia-vasculature modeling. My research background spans metabolomics,
clinical research, and bioinformatics, with a proven track record in developing reproducible workflows, FAIR-compliant data pipelines, and integrative
computational models for complex biomedical datasets.
Currently, I serve as a Senior Biomedical Data Scientist (FAIR Data and Open Science) in the Data Science for Biomedical Research Unit (DSBU) at the University of Lausanne (UNIL),
where I support large-scale data integration and research data management across life sciences and clinical projects. I am passionate about building tools that make scientific data more accessible and
reusable—including developing open-source Python packages for biomedical metadata extraction and standardization.
My expertise bridges clinical research, computational biology, and software development, with proficiency in Python, and version control systems.
I am particularly interested in creating solutions that help researchers adhere to FAIR principles, enhance reproducibility, and accelerate discovery through better data management practices.
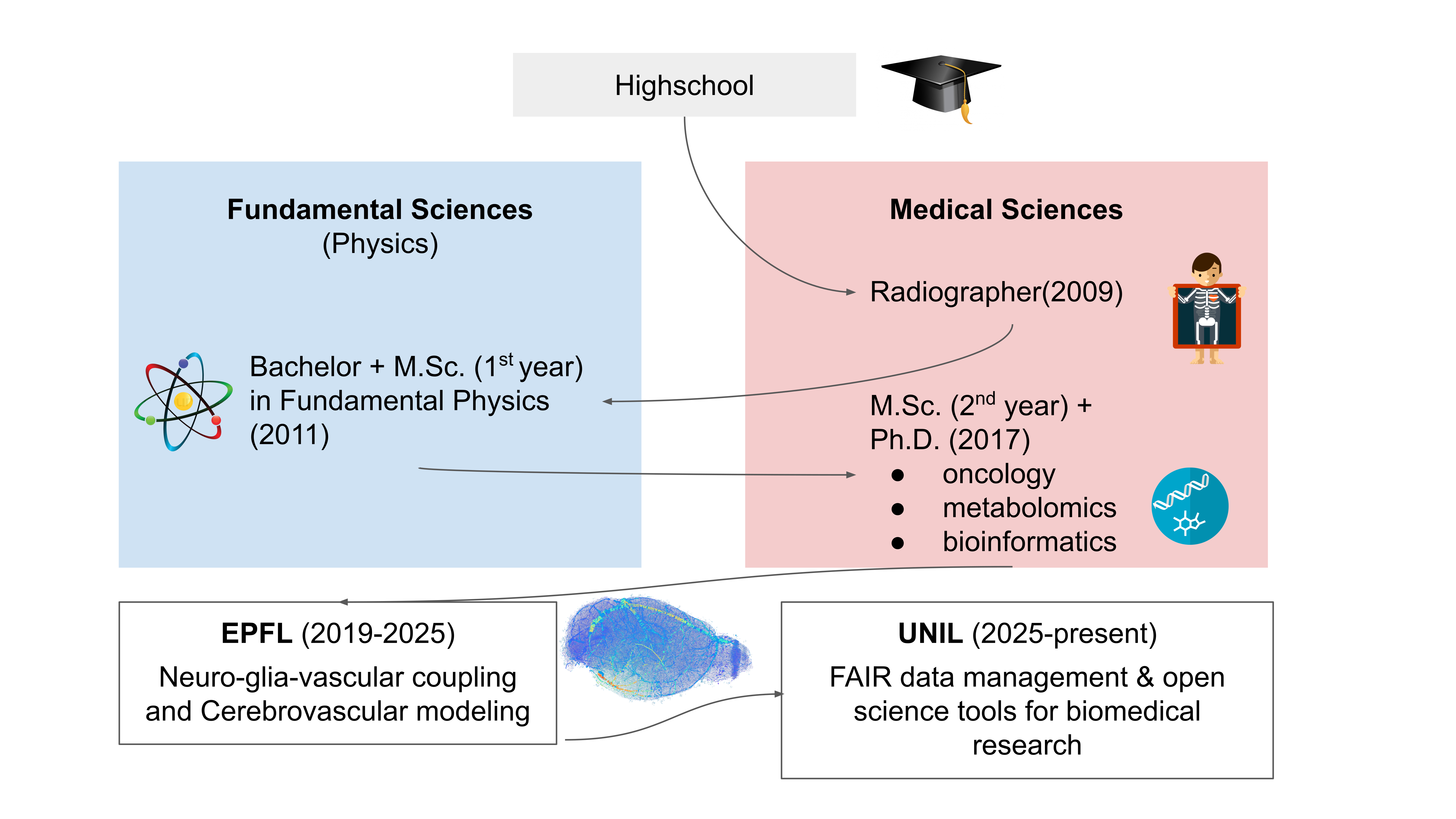
Education and Career
For the full list of skills and experience, see or download the
CV.
Research
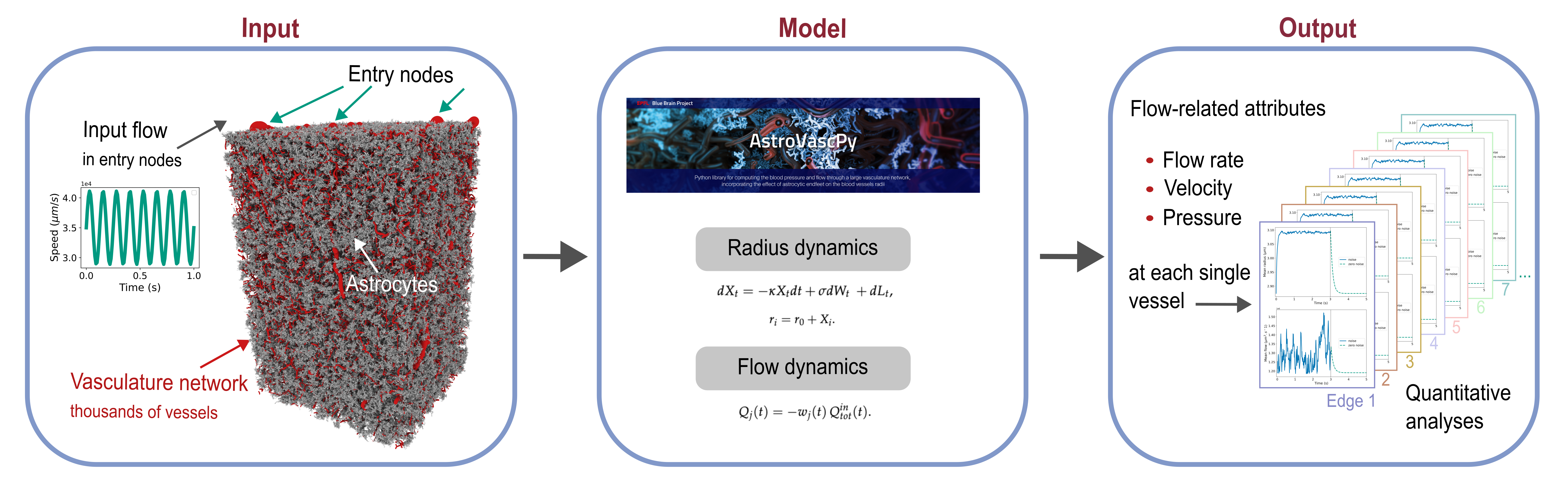
Cerebrovascular simulation: fluid dynamics & astrocyte coupling
How do brain cells control blood supply? This study reveals how astrocytes—the brain's support
cells—regulate blood flow through localized vessel diameter changes. Our computational model shows capillaries
as key players in cerebral perfusion, with layer-specific responses across cortical depths.
[Battini et al., Biomedicines, 2025]
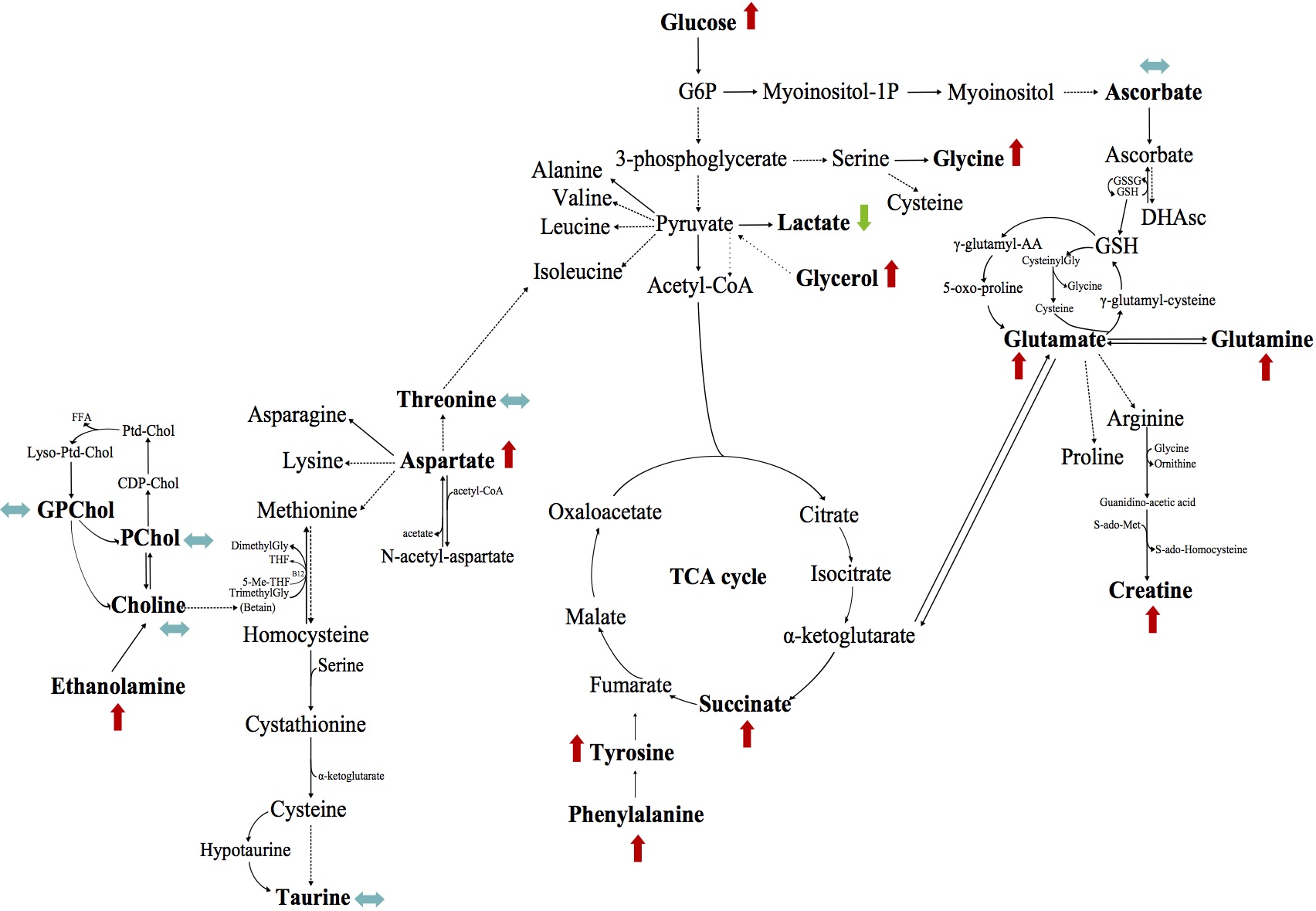
Metabolomics in pancreatic cancer: predicting clinical outcomes
Identified metabolic biomarkers to predict survival in pancreatic cancer patients using
HRMAS NMR spectroscopy. Analyzed 106 tumor samples and discovered that ethanolamine levels distinguish
long-term from short-term survivors—a potential clinical biomarker obtainable in 20 minutes during surgery.
This publication demonstrates how metabolomics profiling can guide treatment decisions.
[Battini et al., BMC Medicine, 2017]
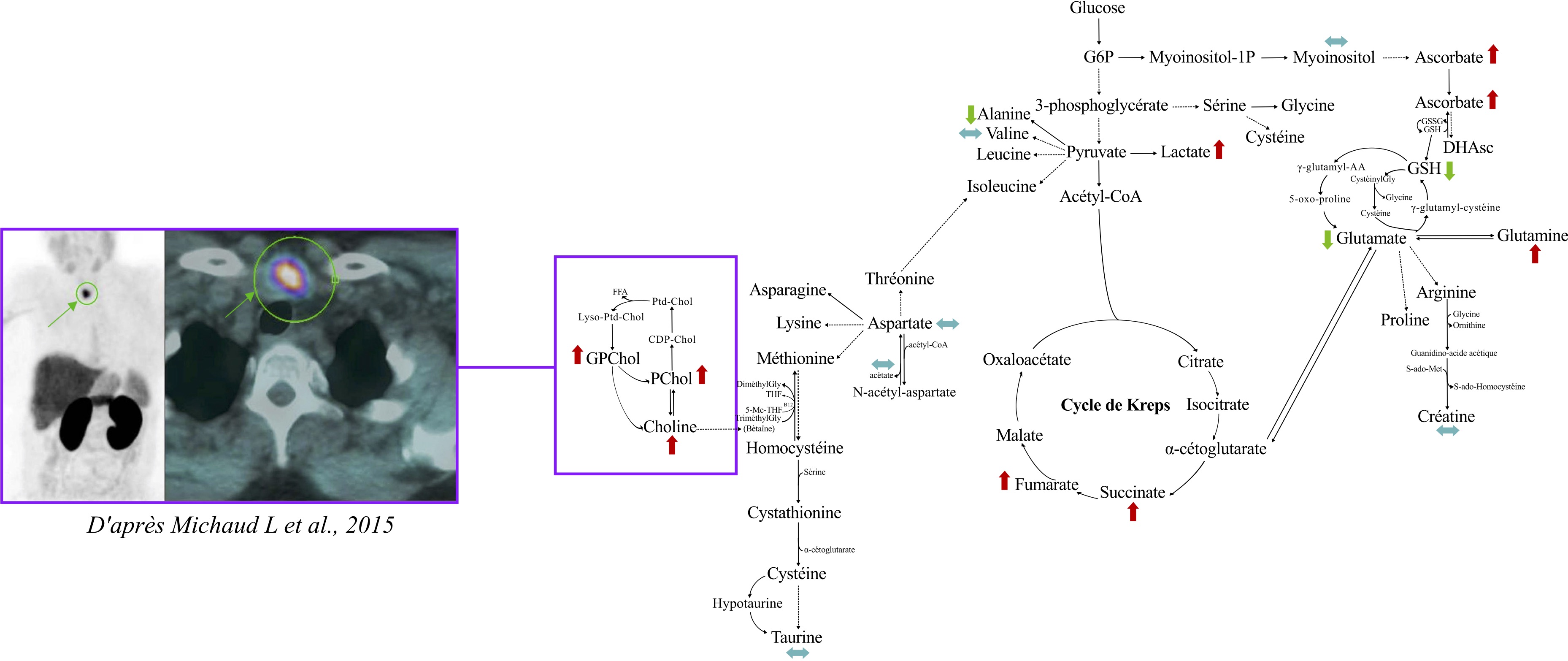
Metabolomics of hyperfunctioning parathyroid glands
Distinguished single-gland from multiglandular disease in primary hyperparathyroidism using HRMAS NMR metabolomics.
Analyzed 43 samples from 32 patients, identifying a 9-metabolite signature (including phosphorylcholine, choline, and glucose)
that accurately differentiates disease subtypes. Applied network analysis with the ADEMA algorithm for metabolite prediction,
offering potential for intraoperative surgical guidance.
[Battini et al., Surgery, 2016]
Projects
AstroVascPy
Python · Computational Neuroscience · HPC
A Python library for multiscale neuro–glia–vascular blood flow modelling, developed at EPFL’s Blue Brain Project.
Focus on graph-based vasculature representations, flow simulations, and integration with existing neuroscience tools.
View on GitHub
FAIR Metadata Pipeline
Python · Snakemake · FAIR · Metadata Standards
Automated Snakemake-based pipeline for harmonizing clinical and research metadata across multiple data sources.
Implements FAIR principles for improved data interoperability and reusability in biomedical research.
In Development
Biomedical Data Validation Suite
Python · PHP · Data Quality · Archival Standards
Comprehensive toolkit for validating, extracting, and packaging clinical research data for long-term archiving.
Ensures compliance with institutional data management standards and facilitates reproducible research workflows.
In Development
Vascular Network Analysis
Python · Graph Theory · Computational Neuroscience
Advanced graph-based analysis tools for modeling cerebrovascular blood flow dynamics.
Combines time-series analysis with machine learning approaches to study neuro-vascular coupling mechanisms.
In Development
Skills
Programming & Data
Python (NumPy, pandas, SciPy, scikit-learn, PyTorch, Matplotlib, Seaborn), R, Bash, LaTeX.
Reproducible workflows with Snakemake, containers (Docker, Singularity), Git/GitHub/GitLab.
Data Management & FAIR
FAIR data principles, metadata standards, data stewardship, REDCap, clinical and research data registries,
Agile project management (Scrum, Kanban), JIRA, Confluence, Trello, Slack.
Selected Publications
-
Modeling of Blood Flow Dynamics in Rat Somatosensory Cortex
Battini S, Cantarutti N, Kotsalos C, et al. Biomedicines, 2025.
Read article
-
Metabolomics approaches in pancreatic adenocarcinoma: tumor metabolism profiling predicts clinical outcome of patients
Battini S, Faitot F, Imperiale A, et al. BMC Medicine, 2017.
Read article
-
High-resolution magic angle spinning 1H nuclear magnetic resonance spectroscopy metabolomics of hyperfunctioning parathyroid glands
Battini S, Imperiale A, Taieb D, et al. Surgery, 2016.
Read article
For the full list of publications, see
Google Scholar.